Regardless of their stage or type, cancers appear to share a telltale signature of widespread changes to the so-called epigenome, according to a team of researchers. In a study published online in Genome Medicine on Aug. 26, the investigators say they have found widespread and distinctive changes in a broad variety of cancers to chemical marks known as methyl groups attached to DNA, which help govern whether genes are turned “on” or “off,” and ultimately how the cell behaves. Such reversible chemical marks on DNA are known as epigenetic, and together they make up the epigenome.
“Regardless of the type of solid tumor, the pattern of methylation is much different on the genomes of cancerous cells than in healthy cells,” says Andrew Feinberg, M.D., M.P.H., a professor of medicine, molecular biology and genetics, oncology, and biostatistics at the Johns Hopkins University School of Medicine. Feinberg led the new study along with Rafael Irizarry, Ph.D., a professor of biostatics at Harvard University and the Dana-Farber Cancer Institute. “These changes happen very early in tumor formation, and we think they enable tumor cells to adapt to changes in their environment and thrive by quickly turning their genes on or off,” Feinberg says.
Feinberg, along with Johns Hopkins University School of Medicine oncology professor Bert Vogelstein, M.D., first identified abnormal methylation in some cancers in 1983. Since then, Feinberg’s and other research groups have found other cancer-associated changes in epigenetic marks. But only recently, says Feinberg, did researchers gain the tools needed to find out just how widespread these changes are.
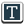
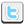
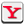
For their study, the research team took DNA samples from breast, colon, lung, thyroid and pancreas tumors, and from healthy tissue, and analyzed methylation patterns on the DNA. “All of the tumors had big blocks of DNA where the methylation was randomized in cancer, leading to loss of methylation over big chunks and gain of methylation in smaller regions,” says Winston Timp, Ph.D., an assistant professor of biomedical engineering at Johns Hopkins. “The changes arise early in cancer development, suggesting that they could conspire with genetic mutations to aid cancer development,” he says.
The overall effect, Feinberg says, appears to be that cancers can easily turn genes “on” or “off” as needed. For example, they often switch off genes that cause dangerous cells to self-destruct while switching on genes that are normally only used very early in development and that enable cancers to spread and invade healthy tissue. “They have a toolbox that their healthy neighbors lack, and that gives them a competitive advantage,” Feinberg says.
“These insights into the cancer epigenome could provide a foundation for development of early screening or preventive treatment for cancer,” Timp says, suggesting that the distinctive methylation “fingerprint” could potentially be used to tell early-stage cancers apart from other, harmless growths. Even better, he says, would be to find a way to prevent the transition to a cancerous fingerprint from happening at all.
Thanks to Johns Hopkins Medicine for contributing this story.